Darin Christensen and Thiemo Fetzer
tl;dr: Fast computation of standard errors that allows for serial and spatial auto-correlation.
Economists and political scientists often employ panel data that track units (e.g., firms or villages) over time. When estimating regression models using such data, we often need to be concerned about two forms of auto-correlation: serial (within units over time) and spatial (across nearby units). As Cameron and Miller (2013) note in their excellent guide to cluster-robust inference, failure to account for such dependence can lead to incorrect conclusions: “[f]ailure to control for within-cluster error correlation can lead to very misleadingly small standard errors…†(p. 4).
Conley (1999, 2008) develops one commonly employed solution. His approach allows for serial correlation over all (or a specified number of) time periods, as well as spatial correlation among units that fall within a certain distance of each other. For example, we can account for correlated disturbances within a particular village over time, as well as between that village and every other village within one hundred kilometers. As with serial correlation, spatial correlation can be positive or negative. It can be made visually obvious by plotting, for example, residuals after removing location fixed effects.
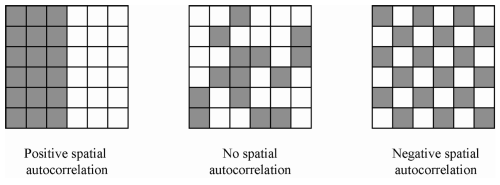
We provide a new function that allows R users to more easily estimate these corrected standard errors. (Solomon Hsiang (2010) provides code for STATA, which we used to test our estimates and benchmark speed.) Moreover using the excellent lfe, Rcpp, and RcppArmadillo packages (and Tony Fischetti’s Haversine distance function), our function is roughly 20 times faster than the STATA equivalent and can scale to handle panels with more units. (We have used it on panel data with over 100,000 units observed over 6 years.)
This demonstration employs data from Fetzer (2014), who uses a panel of U.S. counties from 1999-2012. The data and code can be downloaded here.
STATA Code:
We first use Hsiang’s STATA code to compute the corrected standard errors (spatHAC in the output below). This routine takes just over 25 seconds.
cd "~/Dropbox/ConleySEs/Data"
clear; use "new_testspatial.dta"
tab year, gen(yy_)
tab FIPS, gen(FIPS_)
timer on 1
ols_spatial_HAC EmpClean00 HDD yy_*FIPS_2-FIPS_362, lat(lat ) lon(lon ) t(year) p(FIPS) dist(500) lag(5) bartlett disp
# *-----------------------------------------------
# * Variable | OLS spatial spatHAC
# *-------------+---------------------------------
# * HDD | -0.669 -0.669 -0.669
# * | 0.608 0.786 0.838
timer off 1
timer list 1
# 1: 24.8 / 3 = 8.2650R Code:
Using the same data and options as the STATA code, we then estimate the adjusted standard errors using our new R function. This requires us to first estimate our regression model using the felm function from the lfe package.
# Loading sample data:
dta_file <- "~/Dropbox/ConleySEs/Data/new_testspatial.dta"
DTA <-data.table(read.dta(dta_file))
setnames(DTA, c("latitude", "longitude"), c("lat", "lon"))
# Loading R function to compute Conley SEs:
source("~/Dropbox/ConleySEs/ConleySEs_17June2015.R")
ptm <-proc.time()
# We use the felm() from the lfe package to estimate model with year and county fixed effects.
# Two important points:
# (1) We specify our latitude and longitude coordinates as the cluster variables, so that they are included in the output (m).
# (2) We specify keepCx = TRUE, so that the centered data is included in the output (m).
m <-felm(EmpClean00 ~HDD -1 |year +FIPS |0 |lat +lon,
data = DTA[!is.na(EmpClean00)], keepCX = TRUE)
coefficients(m) %>%round(3) # Same as the STATA result. HDD
-0.669 We then feed this model to our function, as well as the cross-sectional unit (county FIPS codes), time unit (year), geo-coordinates (lat and lon), the cutoff for serial correlation (5 years), the cutoff for spatial correlation (500 km), and the number of cores to use.
SE <-ConleySEs(reg = m,
unit = "FIPS",
time = "year",
lat = "lat", lon = "lon",
dist_fn = "SH", dist_cutoff = 500,
lag_cutoff = 5,
cores = 1,
verbose = FALSE)
sapply(SE, sqrt) %>%round(3) # Same as the STATA results. OLS Spatial Spatial_HAC
0.608 0.786 0.837 proc.time() -ptm user system elapsed
1.619 0.055 1.844 Estimating the model and computing the standard errors requires just over 1 second, making it over 20 times faster than the comparable STATA routine.
R Using Multiple Cores:
Even with a single core, we realize significant speed improvements. However, the gains are even more dramatic when we employ multiple cores. Using 4 cores, we can cut the estimation of the standard errors down to around 0.4 seconds. (These replications employ the Haversine distance formula, which is more time-consuming to compute.)
pkgs <-c("rbenchmark", "lineprof")
invisible(sapply(pkgs, require, character.only = TRUE))
bmark <-benchmark(replications = 25,
columns = c('replications','elapsed','relative'),
ConleySEs(reg = m,
unit = "FIPS", time = "year", lat = "lat", lon = "lon",
dist_fn = "Haversine", lag_cutoff = 5, cores = 1, verbose = FALSE),
ConleySEs(reg = m,
unit = "FIPS", time = "year", lat = "lat", lon = "lon",
dist_fn = "Haversine", lag_cutoff = 5, cores = 2, verbose = FALSE),
ConleySEs(reg = m,
unit = "FIPS", time = "year", lat = "lat", lon = "lon",
dist_fn = "Haversine", lag_cutoff = 5, cores = 4, verbose = FALSE))
bmark %>%mutate(avg_eplased = elapsed /replications, cores = c(1, 2, 4)) replications elapsed relative avg_eplased cores
1 25 23.48 2.095 0.9390 1
2 25 15.62 1.394 0.6249 2
3 25 11.21 1.000 0.4483 4Given the prevalence of panel data that exhibits both serial and spatial dependence, we hope this function will be a useful tool for applied econometricians working in R.
Feedback Appreciated: Memory vs. Speed Tradeoff
This was Darin’s first foray into C++, so we welcome feedback on how to improve the code. In particular, we would appreciate thoughts on how to overcome a memory vs. speed tradeoff we encountered. (You can email Darin at darinc[at]stanford.edu.)
The most computationally intensive chunk of our code computes the distance from each unit to every other unit. To cut down on the number of distance calculations, we can fill the upper triangle of the distance matrix and then copy it to the lower triangle. With [math]N[/math] units, this requires only  [math](N (N-1) /2)[/math] distance calculations.
However, as the number of units grows, this distance matrix becomes too large to store in memory, especially when executing the code in parallel. (We tried to use a sparse matrix, but this was extremely slow to fill.) To overcome this memory issue, we can avoid constructing a distance matrix altogether. Instead, for each unit, we compute the vector of distances from that unit to every other unit. We then only need to store that vector in memory. While that cuts down on memory use, it requires us to make twice as many   [math](N (N-1))[/math]  distance calculations.
As the number of units grows, we are forced to perform more duplicate distance calculations to avoid memory constraints – an unfortunate tradeoff. (See the functions XeeXhC and XeeXhC_Lg in ConleySE.cpp.)
sessionInfo()R version 3.2.2 (2015-08-14)
Platform: x86_64-apple-darwin13.4.0 (64-bit)
Running under: OS X 10.10.4 (Yosemite)
locale:
[1] en_GB.UTF-8/en_GB.UTF-8/en_GB.UTF-8/C/en_GB.UTF-8/en_GB.UTF-8
attached base packages:
[1] stats graphics grDevices utils datasets methods
[7] base
other attached packages:
[1] RcppArmadillo_0.5.400.2.0 Rcpp_0.12.0
[3] geosphere_1.4-3 sp_1.1-1
[5] lfe_2.3-1709 Matrix_1.2-2
[7] ggplot2_1.0.1 foreign_0.8-65
[9] data.table_1.9.4 dplyr_0.4.2
[11] knitr_1.11
loaded via a namespace (and not attached):
[1] Formula_1.2-1 magrittr_1.5 MASS_7.3-43
[4] munsell_0.4.2 xtable_1.7-4 lattice_0.20-33
[7] colorspace_1.2-6 R6_2.1.1 stringr_1.0.0
[10] plyr_1.8.3 tools_3.2.2 parallel_3.2.2
[13] grid_3.2.2 gtable_0.1.2 DBI_0.3.1
[16] htmltools_0.2.6 yaml_2.1.13 assertthat_0.1
[19] digest_0.6.8 reshape2_1.4.1 formatR_1.2
[22] evaluate_0.7.2 rmarkdown_0.8 stringi_0.5-5
[25] compiler_3.2.2 scales_0.2.5 chron_2.3-47
[28] proto_0.3-10